Dfind old genome assemblies8/2/2023  SyRI provides an efficient method for accurate identification of all common rearrangements including transpositions and duplications. Moreover, commonly used tools have limited functionality in identifying transpositions (i.e., the relocation of a sequence within a chromosome) and distal duplications. This is a significant improvement compared to current methods which typically do not predict both breakpoints for all rearrangements in both of the genomes. SyRI annotates the coordinates of rearranged regions (i.e., breakpoints on both sides of a rearrangement in both genomes) providing a complete regional annotation of rearrangements. Afterwards, SyRI identifies local sequence differences within both the rearranged and the non-rearranged (syntenic) regions. SyRI expects whole-genome alignments (WGA) as input and starts by searching for differences in the structures of the genomes. Here, we introduce SyRI (Synteny and Rearrangement Identifier), a method to identify structural as well as sequence differences between two whole-genome assemblies. Available tools include AsmVar, which compares individual contigs of an assembly against a reference sequence and analyzes alignment breakpoints to identify inversions and translocations Assemblytics, which utilizes uniquely aligned regions within contig alignments to a reference sequence to identify various types of genomic differences including large indels or differences in local repeats and Smartie-sv, which compares individual alignments between assembly and reference sequences. However, despite recent technological improvements to simplify the generation of whole-genome de novo assemblies, there are so far only a few tools which use whole-genome assemblies as the basis for the identification of genomic differences. In contrast, whole-genome assemblies enable the identification of complex rearrangements as the assembled contigs are typically much longer and of higher quality as compared to raw sequence reads. Even though such alignments allow to find local sequence differences (like SNPs, indels, and structural variations) with high accuracy, accurate prediction of structural differences remains challenging. Many of the state-of-the-art methods used to predict genomic differences utilize short or long read alignments against reference sequences. But even though this information is usually not considered when analyzing whole-genome sequencing data, differences in genome structure are relevant as they can be the basis for diseases phenotypes, reproductive strategies, and survival strategies. Like translocations, inversions and duplications also introduce differences in the genome structure by changing location, orientation, and/or copy number of specific sequences. Although such a translocation could be described as a deletion at one region and an insertion at the other region, this annotation would miss the information that the deleted/inserted sequence is the same and that the deleted sequence is actually not deleted but rather relocated to a different region. 
For example, a translocation is a genomic rearrangement where a specific sequence has moved from one region in the genome to another region. 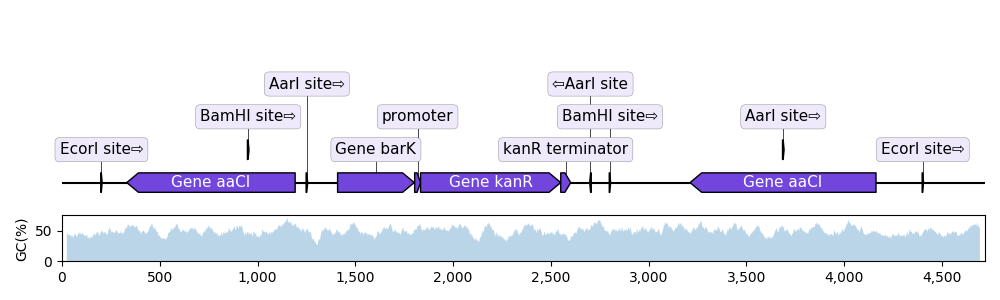
But even though the annotation of all sequence differences against a reference sequence would be sufficient to reconstruct the actual sequence of a genome, sequence differences alone cannot describe the complex genomic rearrangements. Differences in genomes can range from single nucleotide differences to highly complex genomic rearrangements, and they are commonly described as local sequence differences in comparison to a reference sequence. Genomic differences form the basis for phenotypic variation and allow us to decipher evolutionary past and gene function. 
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
